IPython, Jupyter Notebooks, and matplotlib
Two and a half years ago, I wrote a post about programming in Python. One of my tips was to use the Python shell, so that one can quickly test simple commands before integrating them in a more complicated project.
Fast forward until now, and my Python habits have changed substantially. One notable change I have made is to use IPython instead of the Python shell. For my usage purposes, the IPython shell has been a strictly superior version of the standard one due to the following:
- It includes TAB completion for functions. For instance, suppose I’m importing the numpy library,
and I want to create an array variable, which means I need the
arrayfunction. I start the IPython shell (by typingipythonon the command line), import the numpy library, and when I press the TAB key aftera = np.arr, I get the output:
In [1]: import numpy as np
In [2]: a = np.arr
np.array np.array2string np.array_equal np.array_equiv np.array_repr np.array_split np.array_str IPython is smart enough to tell me which methods I might be interested in using! It’s a really nice feature, and I’ve found that it also works when one tries to autocomplete function parameters. In the standard Python shell, typing TAB just means … creating extra TABs.
- It makes it easier to fix for loops, which is handy because it’s really easy to make a mistake with loops. Consider the trivial example below:
In [2]: for i in range(4):
...: print "hi""
...:
File "<ipython-input-2-f57d06c46d12>", line 2
print "hi""
^
SyntaxError: EOL while scanning string literalIn IPython, to fix the loop, I just need to press the UP key and it will load both lines of the
for loop. In the standard shell, the UP key would only return print "hi"", forcing the user
to essentially retype the loop.
- It remembers commands from previous sessions, so I can exit an IPython session, do other stuff, then restart IPython, press the UP key, and it will give me the commands I used in my last session.
These three are the extra IPython features that have been most useful for my work.
I frequently use Python for work because it is a simple language that has lots of robust math, machine learning, and data analysis libraries. My favorite Python library is matplotlib, which is used for forming high-quality plots.
A few months ago, my workflow for using matplotlib was to write a script that first gets the data
into a matplotlib plot, and then saves it (using the savefig(...) function). When I need to
make lots of figures, however, it gets cumbersome to manage them, and I often have to keep multiple
images open so I can spot-check their changes when I re-run my script (e.g., if I modified the font
size of the text).
Fortunately, I discovered Jupyter Notebooks. These are brower-based platforms that make managing matplotlib-based images far easier by keeping information unified in one screen.
To start a notebook session, I type in ipython notebook on the command line, which opens up a
web browser (for me, it’s Firefox). I then click New -> Python 2 to start the session. For a basic
plot, I can start by importing the library: import matplotlib.pyplot as plt, but then —
crucially — I use the %matplotlib inline command. The reason for using that is so that when
I write code to plot, and then execute it with a simple SHIFT-ENTER, the image will appear directly
under that code cell. Here’s a simple example:
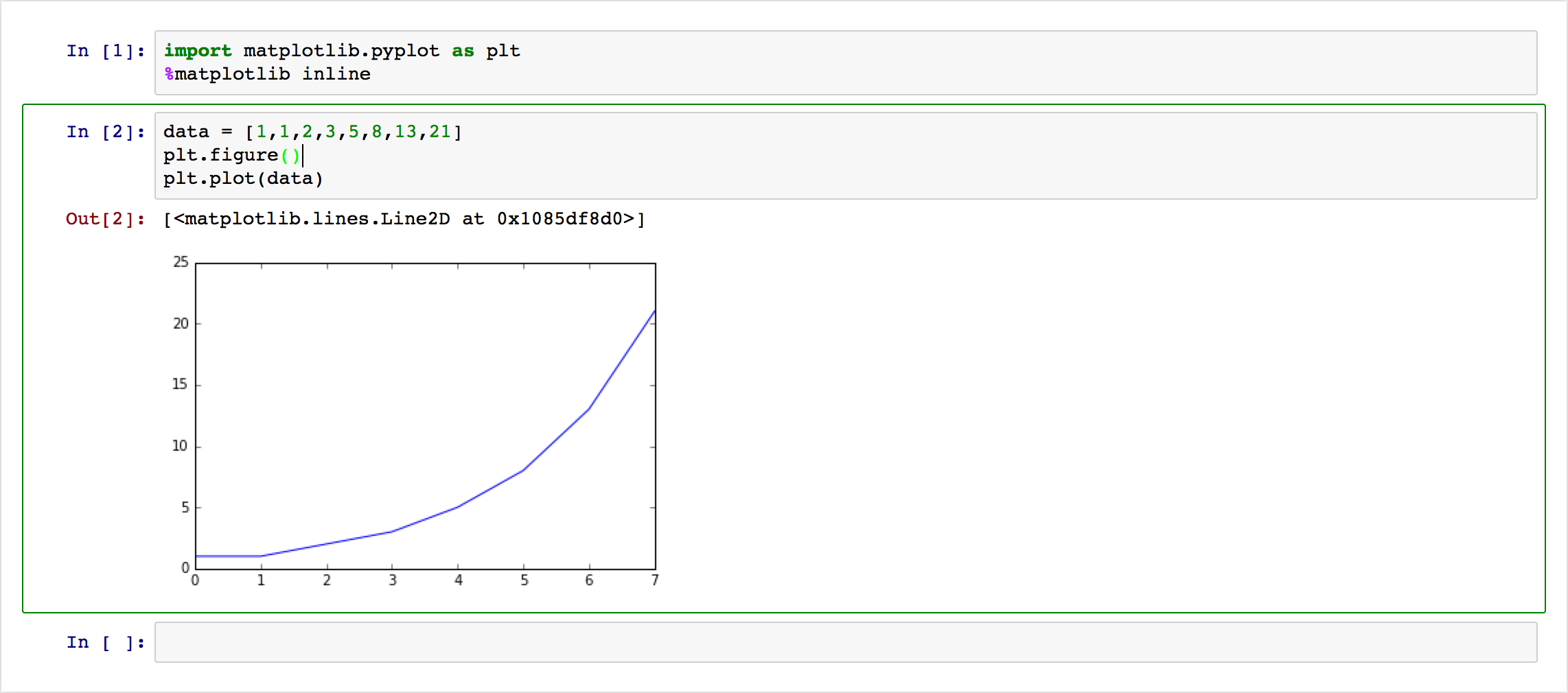
This is nice, but what if I want to change some plot setting? If these images are going to be in an academic paper, they better have labels and legends, among other things. With these notebooks, one can modify the text in a cell and regenerate the image; here’s an example with some common commands I use for my plots:
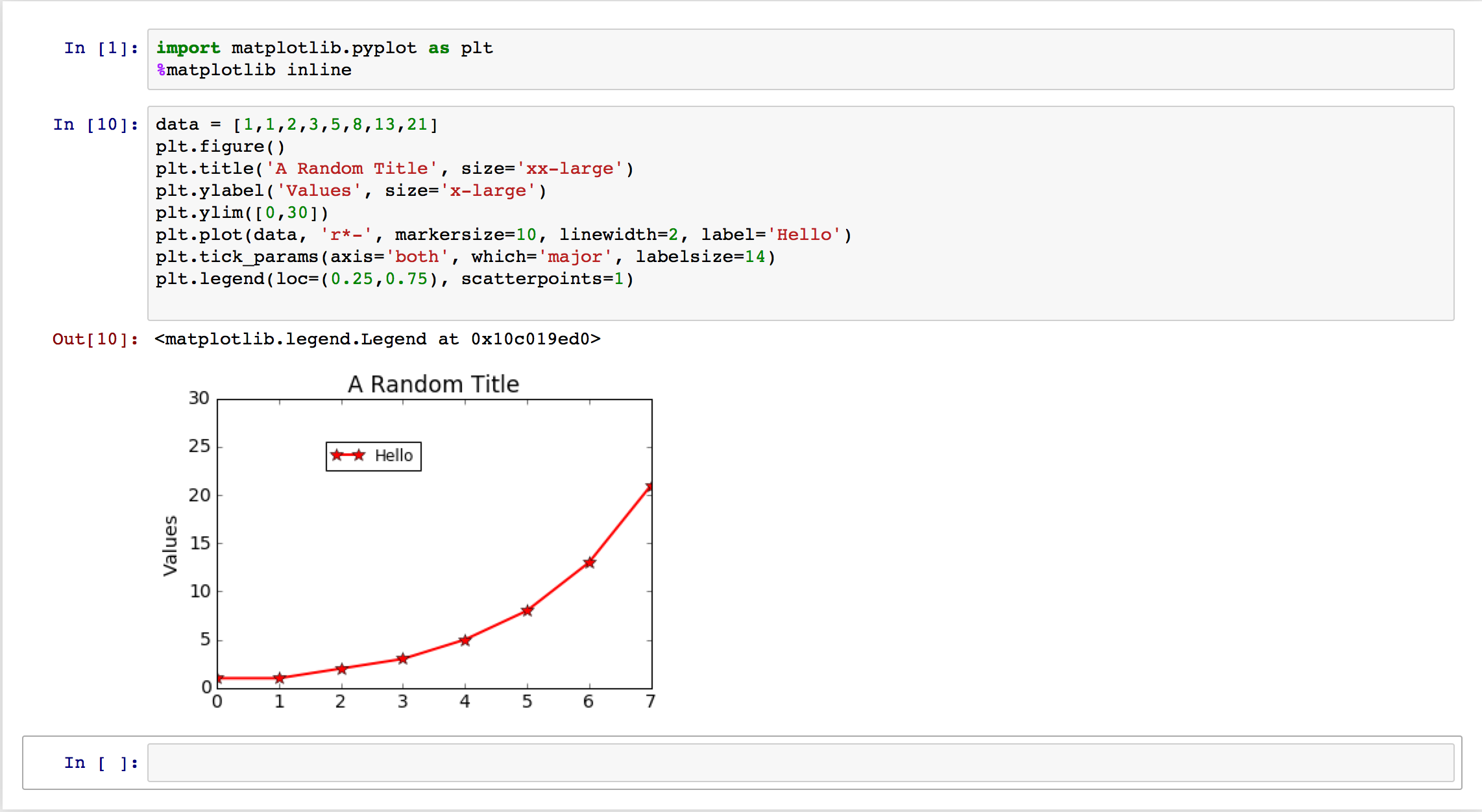
This example doesn’t quite show the benefit enough, but once projects get more complicated, notebooks are a valuable tool to keep data organized. Moreover, one can save a notebook session so that the next time it gets opened again, its plots remain visible on the webpage.
For those of you who use Python, I encourage you to check out IPython and Jupyter. They add on to what is already an awesome general-purpose programming language.